Docking with the AutoDock Suite
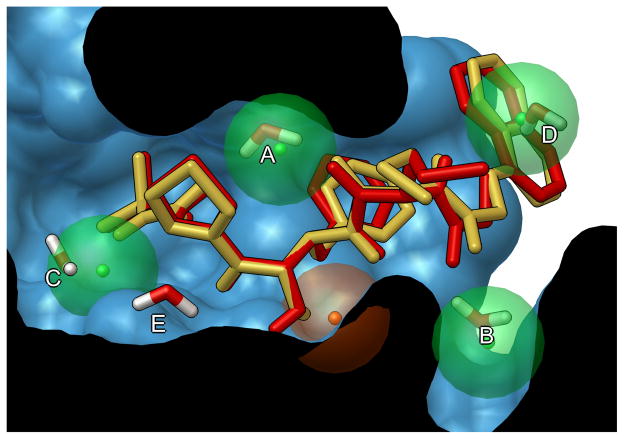
Computational docking is widely used for study of protein-ligand interactions and for drug discovery and development. Typically the process starts with a target of known structure, such as a crystallographic structure of an enzyme of medicinal interest. Docking is then used to predict the bound conformation and binding free energy of small molecules to the target. Single docking experiments are useful for exploring the function of the target, and virtual screening, where a large library of compounds are docked and ranked, may be used to identify new inhibitors for drug development.
AutoDock is a suite of free open–source software for the computational docking and virtual screening of small molecules to macromolecular receptors. The suite currently includes several complementary tools:
Computational Docking Software
Interactive Graphical User Interfaces
Binding Site Prediction
Getting Started
The best way to get started with docking is to follow the detailed tutorial in our Nature Protocols paper: “Computational Ligand-Protein Docking and Virtual Drug Screening with the AutoDock Suite. It is freely available on PubMedCentral, and also available in nice formatted form through Nature Protocols. Be sure to download the supplementary files, which include several worked examples that you can try out. This tutorial will introduce you to three programs with the core functionality of AutoDock:
- AutoDockTools, the graphical user interface you will need for preparing ligand and protein coordinate files, and analyzing results;
- AutoDock Vina, a simple one-step docking program that is effective for most ligand-protein systems;
- AutoDock, with AutoGrid, performs docking in two steps, and provides more user-tunable options for systems with special challenges.
Several specialized docking programs are available as you become more familiar with docking and encounter systems that require special treatment. These include:
- AutoDockFR: a computational docking program with flexible protein targets, including sidechain motion and induced fit;
- AutoDockCrankPep: a program for computational docking of peptides to protein targets.
If you are interested in virtual screening, Raccoon2 provides a dedicated graphical user interface for managing coordinates and docking, and analyzing results.
We have also created specialized tools for different aspects of docking preparation and analysis, such as the binding site prediction software AutoSite and AutoLigand.
Getting Help
ADL, the AutoDock Mailing List, is intended for novice and expert users of the AutoDock suite of programs. It brings together a community of people that have common interests in molecular modeling software related to docking and AutoDock, and are here for learning or helping others. It is also a forum for suggesting new features, and new software versions and functionality will be announced on the list.
Documentation, tutorials, download instructions, and other useful information is available at the sites for each software.