Welcome
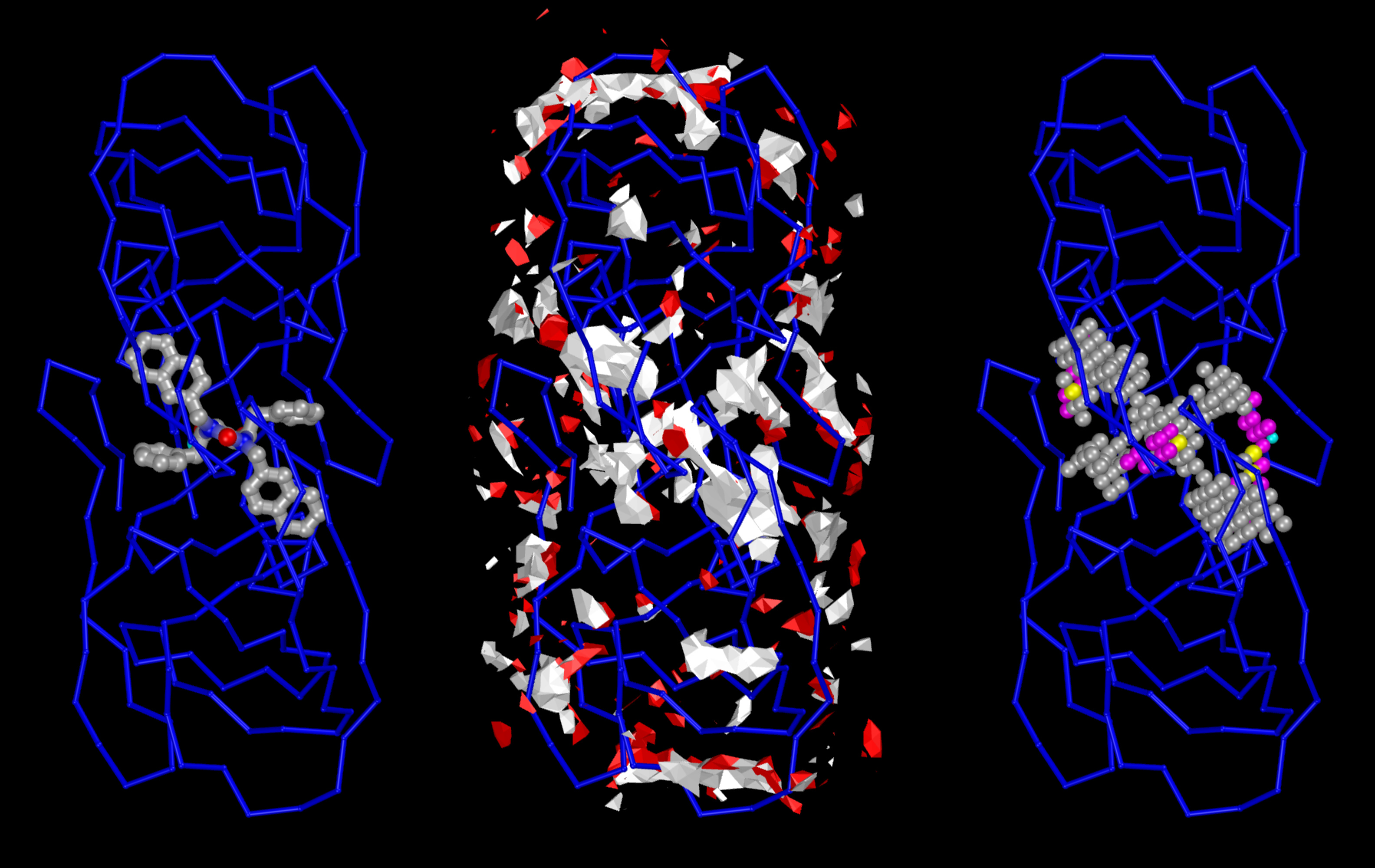
The Center for Computational Structural Biology (CCSB) currently comprises four highly synergistic and independently funded research laboratories focusing on a variety of aspect of modelling and simulating structural biology ranging from biomolecular visualization, computation and design, spanning from atomic structure and interactions to the mesoscale structure of living cells. It was created in 2018 as a successor of the Molecular Graphics Laboratory (MGL), founded by Arthur Olson in 1981. Over the years we have developed several freely-available software tools including AutoDock, Python Molecular Viewer, and CellPACK. Current applications include the design and virtual screening of covalent inhibitors of HIV-1 proteins, docking and binding energy prediction of small molecules and peptides to flexible biomolecular targets, and modeling of subcellular structures such as bacterial nucleoids and secretory vesicles.
Projects
Principal Investigators

Arthur J. Olson, Professor
Arthur Olson has pioneered the use of computer graphics and simulation for the visualization and study of biomolecular structure and function.(faculty website)
Stefano Forli
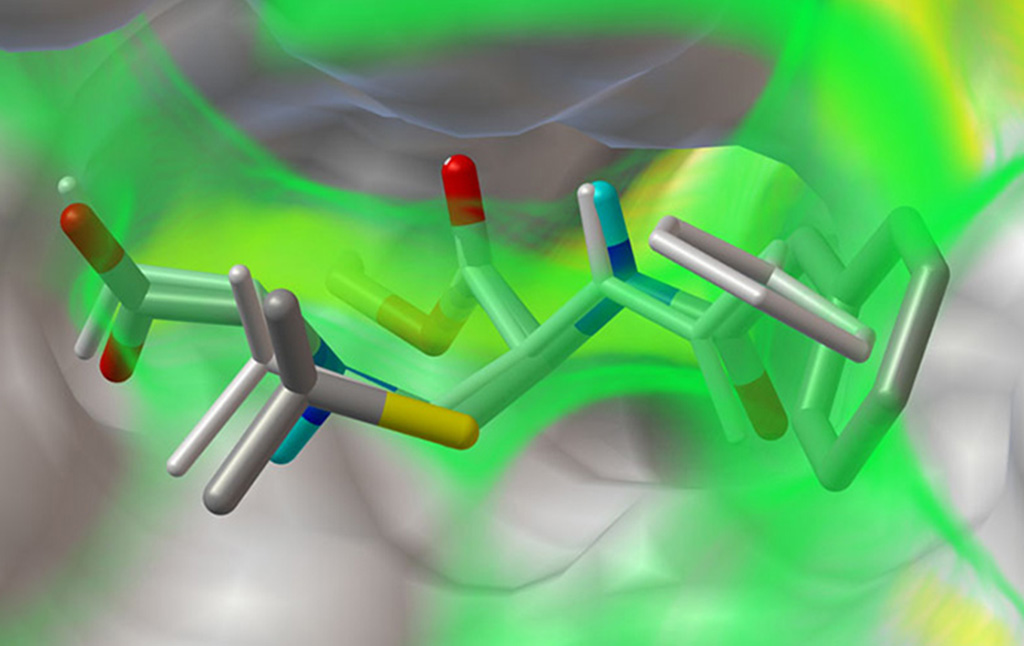
Stefano Forli is developing new methods for computational drug design, discovery, and virtual screening, and applying them to systems of biomedical interest such as HIV-1 infection and cancer. (Forli Lab)

Michel F. Sanner
Michel Sanner has pioneered a strategy for developing interoperable software components for the visualization and modelling of biomolecular interactions. He is also actively pushing the envelop of automated docking with new software such as AutoDockFR and AutoDock CrankPep. (faculty website).

Ludovic Autin
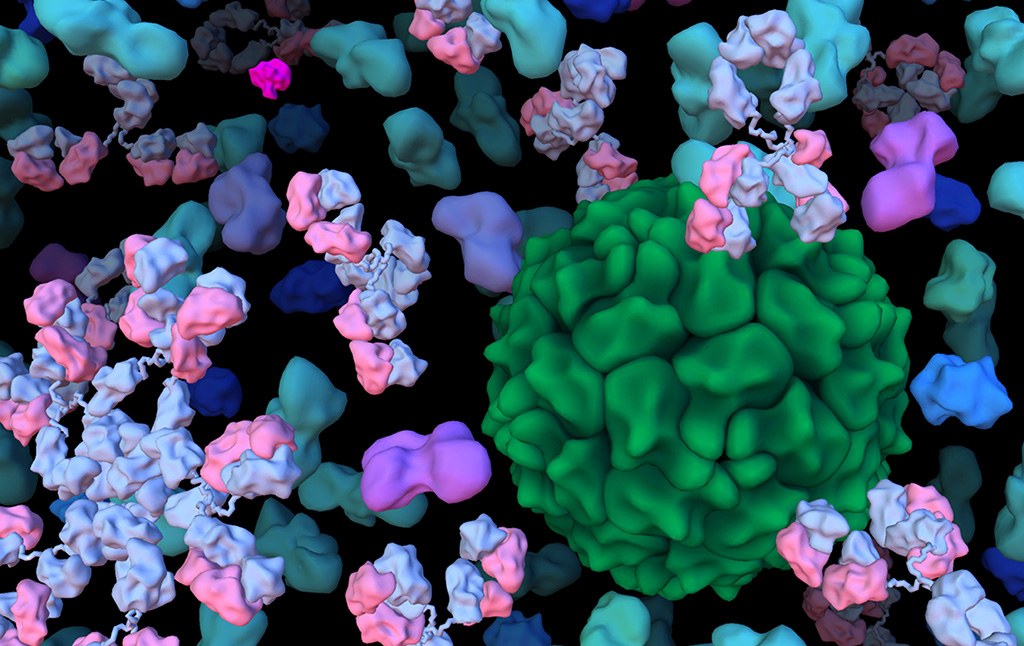
Ludovic’s work focus to fill the gaps existing between scientific visualization and artistic illustration leading to better understanding and a better view of some highly complex scientific data: such as induced-fit molecular recognition, viral unit self-assembly, the complex mechanism of DNA structure, and crowded cellular environments. We are developing software frameworks dedicated to multiscale molecular modeling and visualization: ePMV (protein visualization in third party 3D software), cellPACK (crowded cellular molecular modeling), mesoscope ( web interface for integrating data ), cellPAINT ( illustration and interaction with molecular landscape).
News & Announcements
David S. Goodsell
David Goodsell has officially retired on January 2025. David has divided his time between computational modeling of the mesoscale structure of cells and science education at the RCSB Protein Data Bank. You can still access his work at his personal website.
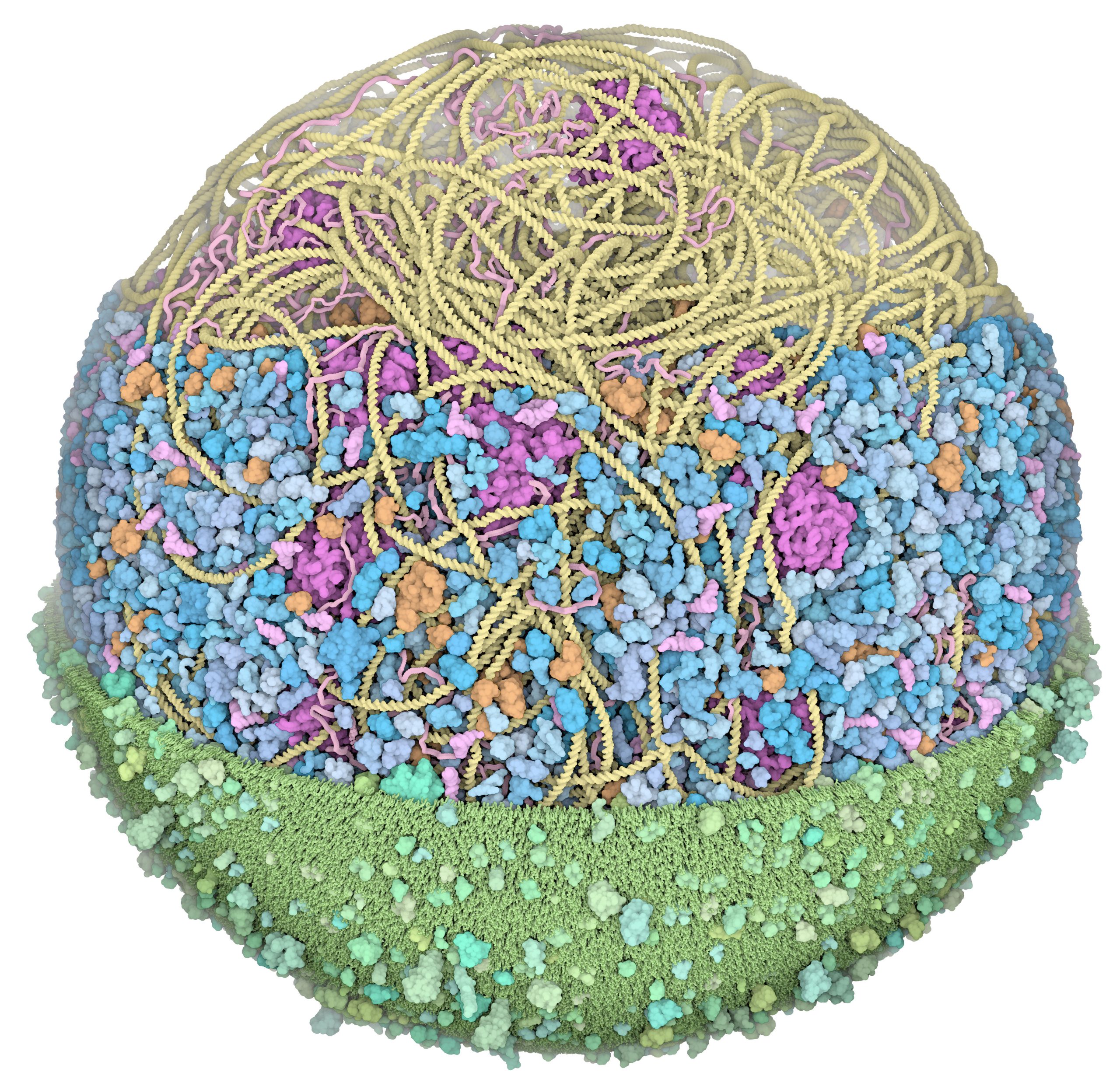
We presented the first structural model of an entire cell in the Journal of Molecular Biology article: ‘Building Structural Models of a Whole Mycoplasma Cell’, https://doi.org/10.1016/j.jmb.2021.167351. Click on the image for a gallery of images of the model, free for use under a Creative Commons license!